LZGraphs¶
LZGraphs is a high-performance Python library for analyzing immune receptor repertoires using Lempel-Ziv 76 compression graphs. Built on a C core, it transforms CDR3 sequences into probabilistic directed graphs that support exact probability computation, constrained sequence generation, and analytical diversity measurement — all without alignment or reference genotypes.
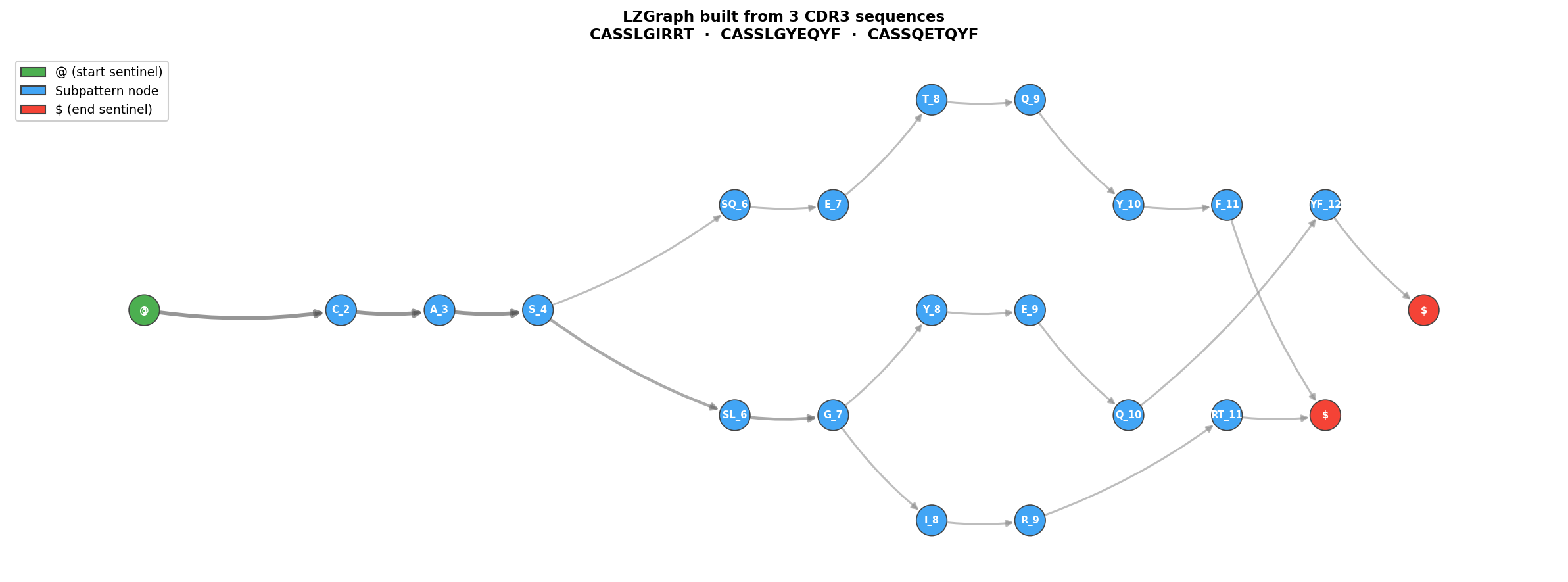
Quick Start¶
from LZGraphs import LZGraph
graph = LZGraph(['CASSLEPSGGTDTQYF', 'CASSDTSGGTDTQYF', 'CASSLEPQTFTDTFFF'],
variant='aap')
graph.lzpgen('CASSLEPSGGTDTQYF') # log generation probability
graph.simulate(1000, seed=42) # generate new sequences
graph.hill_number(2) # inverse Simpson diversity
graph.predicted_richness(100_000) # richness at sequencing depth
lzg build repertoire.tsv -o rep.lzg # build from the command line
lzg diversity rep.lzg # diversity report
lzg simulate rep.lzg -n 10000 > synth.txt # generate sequences
What LZGraphs does¶
Score sequences¶
Compute the exact generation probability of any CDR3 under the repertoire model with lzpgen().
Generate sequences¶
Simulate novel sequences via LZ-constrained random walks — with optional V/J gene constraints.
Measure diversity¶
Hill numbers, Shannon entropy, predicted richness, sample overlap, and sharing spectra — analytically from the graph.
Compare repertoires¶
Jensen-Shannon divergence, cross-scoring, and graph set operations (union, intersection, difference).
Extract ML features¶
Project repertoires into fixed-size feature vectors for classification, clustering, and regression.
Personalize models¶
Bayesian posterior updates to adapt a population graph to an individual patient.
Documentation¶
Learn Installation, quick start, tutorials, and worked examples.
Guides Task-oriented recipes: data prep, generation, comparison, ML features.
Concepts LZ76 algorithm, probability model, graph variants, distribution analytics.
Reference
Complete API for LZGraph, SimulationResult, CLI tool, and exceptions.
C Performance¶
Build graphs from 5,000 sequences in 80 ms. Simulate at ~5,000 seqs/sec. Save/load in < 1 ms.
LZ76 Constraints¶
Every simulated sequence is a valid LZ76 decomposition. No biologically impossible outputs.
If you use LZGraphs in your research, please cite our paper. GitHub · Issues · Contact

